CryoEM Structure Refinement by Integrating NMR Chemical Shifts
€ 24.00 · 4.6 (640) · Auf Lager


Environmentally Ultrasensitive Fluorine Probe to Resolve Protein Conformational Ensembles by 19F NMR and Cryo-EM

CryoEM Structure Refinement by Integrating NMR Chemical Shifts with Molecular Dynamics Simulations

Structure of the 30 kDa HIV-1 RNA Dimerization Signal by a Hybrid Cryo-EM, NMR, and Molecular Dynamics Approach - ScienceDirect

Integrating model simulation tools and cryo‐electron microscopy - Beton - 2023 - WIREs Computational Molecular Science - Wiley Online Library

Experimental NMR and EM data of the 468 kDa TET2 assembly. a MAS NMR
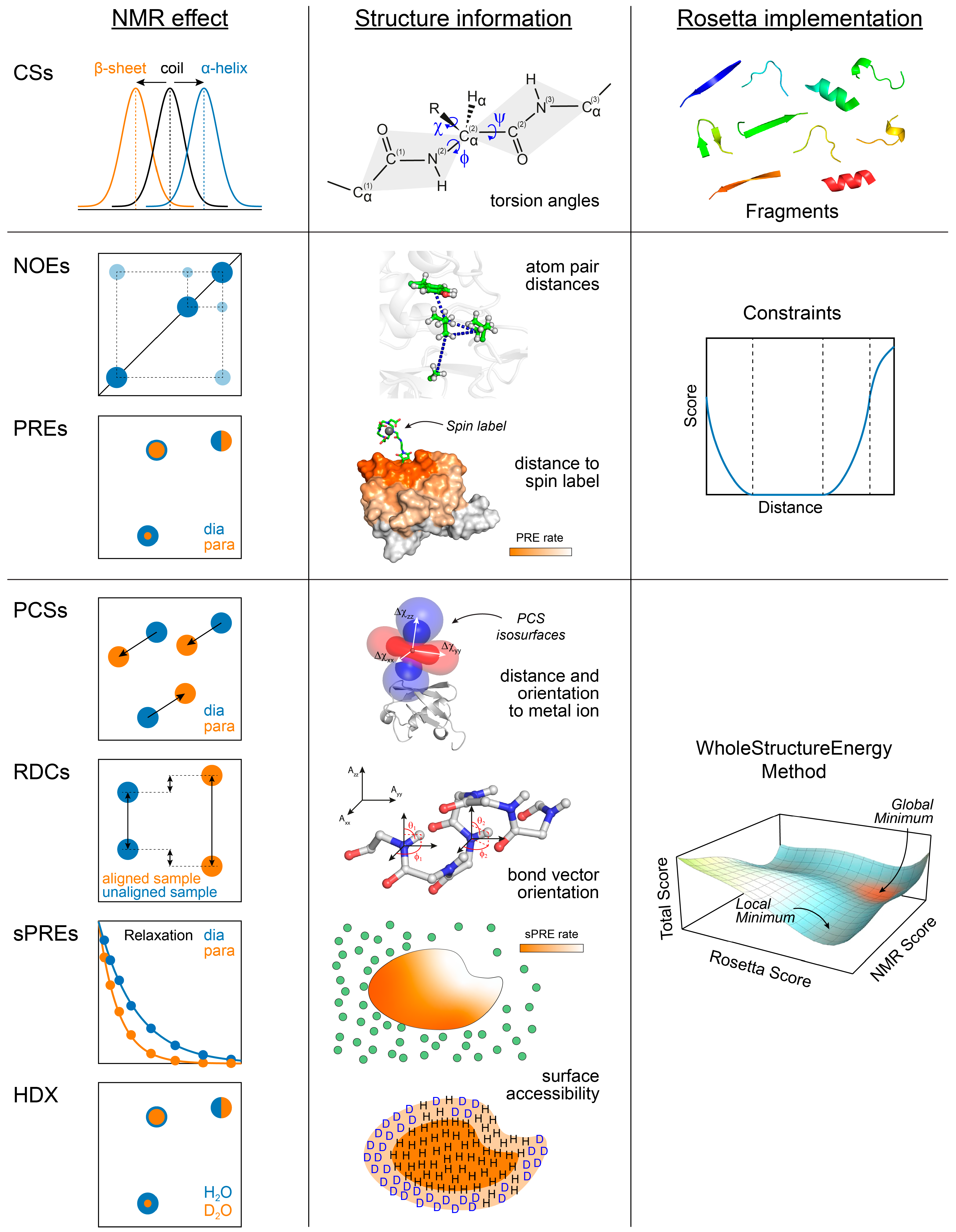
IJMS, Free Full-Text

PDB 5upw structure summary ‹ Protein Data Bank in Europe (PDBe) ‹ EMBL-EBI
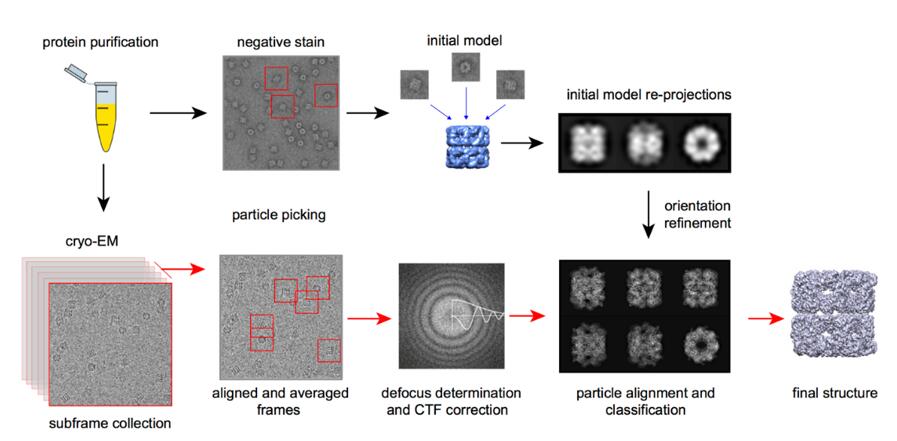
Comparison of Crystallography, NMR and EM - Creative Biostructure
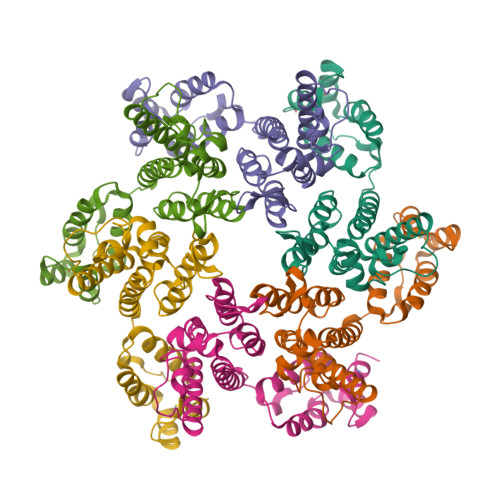
RCSB PDB - 5UPW: CryoEM Structure Refinement by Integrating NMR Chemical Shifts with Molecular Dynamics Simulations

CryoEM Structure Refinement by Integrating NMR Chemical Shifts with Molecular Dynamics Simulations

A) Localization of the dipolar vectors and chemical shift tensors
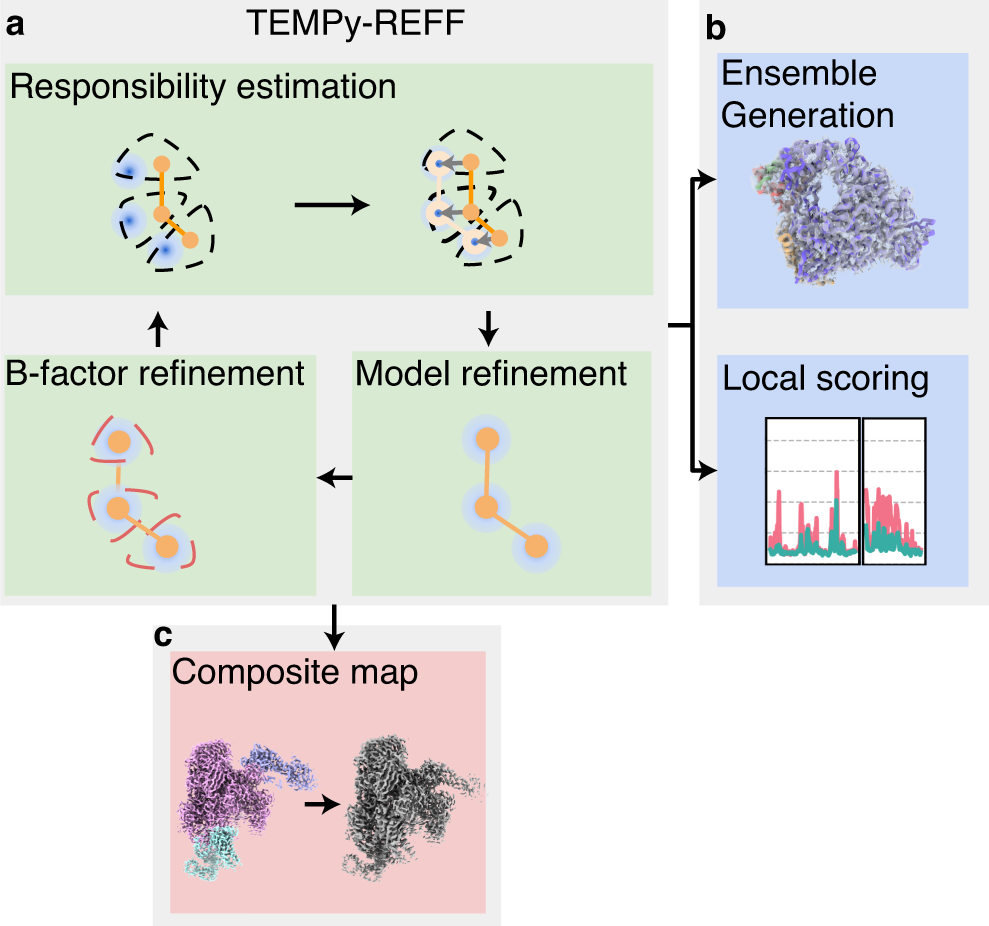
Cryo-EM structure and B-factor refinement with ensemble representation

Imaging active site chemistry and protonation states: NMR crystallography of the tryptophan synthase α-aminoacrylate intermediate
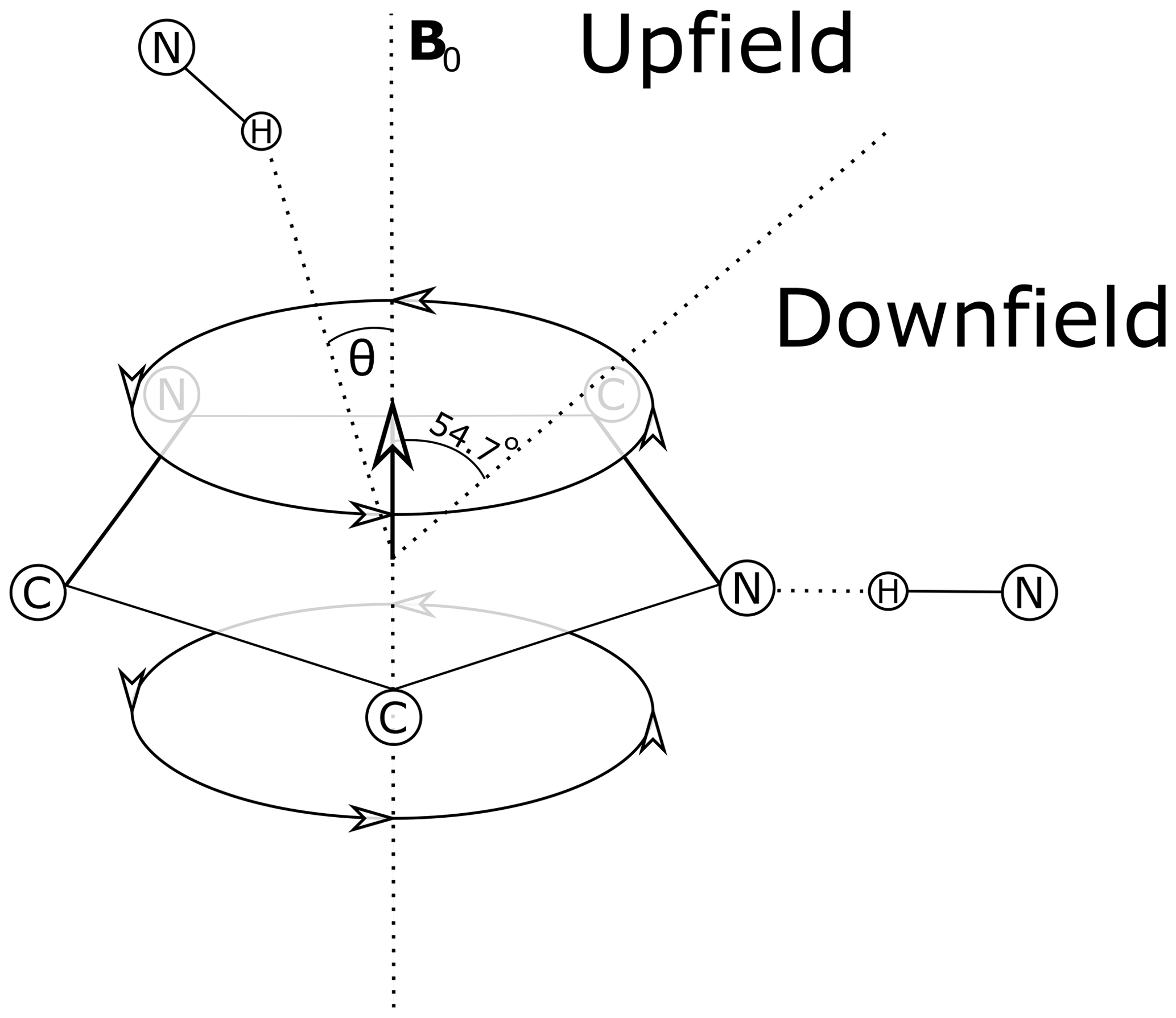
MR - Anomalous amide proton chemical shifts as signatures of hydrogen bonding to aromatic sidechains

Using NMR Chemical Shifts and Cryo-EM Density Restraints in Iterative Rosetta-MD Protein Structure Refinement